
| Homepage | |
| Group Members | |
| Publications | |
| 2002 - | |
| 1988 - 2001 | |
| Theses and | |
| Reports | |
| Picture Gallery | |
| Data | |
| Databases | |
| Software | |
| Job Vacancies | |
| Conferences, | |
| Workshops | |
| and Courses | |
| Local Info | |
| Group Info | |
| Projects | |
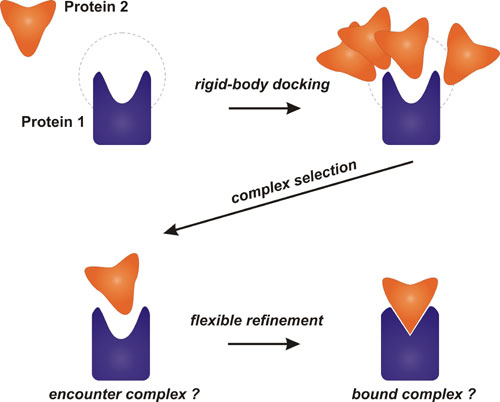 |
A protocol for computational modeling of protein-protein association and protein-protein structure prediction is under development.
The method consists of Rigid Body docking and Flexible Refinement stages. In Rigid Body docking, Brownian Dynamics based sampling is used as implemented in
SDA program.
Available biochemical data is incorporated in the sampling stage. Selected encounter complexes are refined using Molecular Dynamics with implicit solvent model
NPSA.
|
|
|