More
examples of the usage of PIPSA 2.0
(Protein Interaction Property
Similarity Analysis)
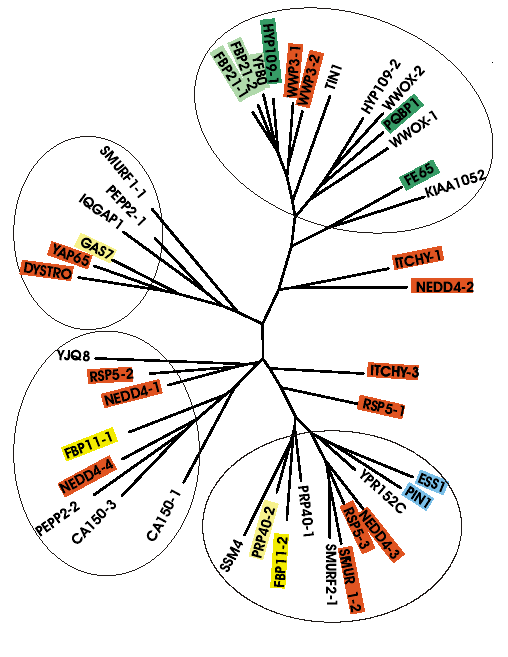
WW domains are small globular protein
interaction
modules found in a wide spectrum of proteins. They recognize their
target
proteins by binding specifically to short linear peptide motifs that
are
often proline-rich. To understand the determinants of the ligand
binding
propensities of WW domains, 42 WW domains were analyzed to derive
quantitative
structure-activity relationships .
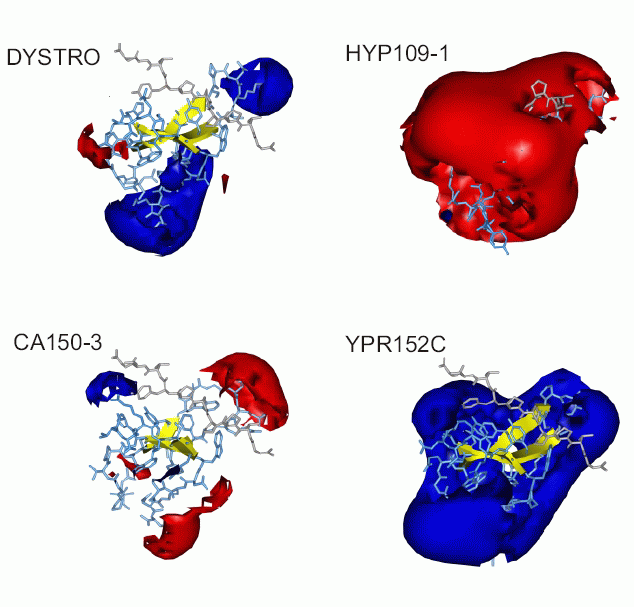
From a protein interaction property
similarity
analysis (PIPSA) of the WW domain structures, a structure-based
classification
of WW domains is proposed that expands the existent
sequence-based
classification scheme.


De Rienzo, F., Gabdoulline,R.R.,
Menziani,M.C.,
De Benedetti, P.G. and Wade,R.C.
Electrostatic Analysis and Brownian
Dynamics Simulation of the Association of Plastocyanin and Cytochrome F
Biophys. J. (2001) 81, 3090-3104.
The oxidation of cytochrome f by the
soluble
cupredoxin plastocyanin is a central reaction in the photosynthetic
electron
transfer chain of all oxygenic organisms. Here, two different
computational
approaches are used to gain new insights into the role of molecular
recognition
and protein-protein association processes in this redox reaction.
A comparative analysis of the computed
molecular electrostatic potentials of seven single and multiple point
mutants
of spinach plastocyanin (D42N, E43K, E43N, E43Q/D44N, E59K/E60Q,
E59K/E60Q/E43N,
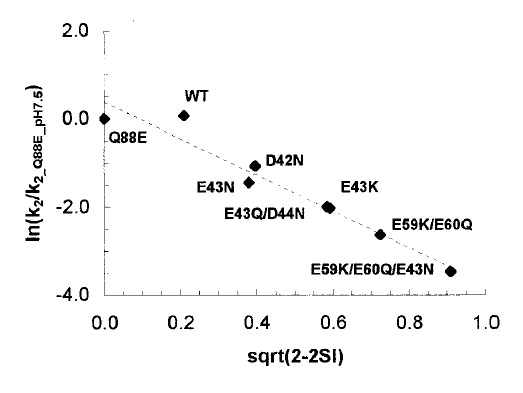
Q88E) and the wt protein was carried out. The experimentally determined
relative rates (k2) for the set of plastocyanin mutants are found to
correlate
well (r2 0.90 0.97) with the computed measure
of
the similarity of the plastocyanin electrostatic potentials:
Razif Gabdoulline, 2005
Back to the index