Nardi,F., Kemmink,J., Sattler,M. and Wade,R.C.
J. Biomol. NMR (2000) 17, 63-77.
| List of PDB structures analysed in the SCAN3D database (specified by the 4-letter code and the subunit identifier): |
| 1iro, 2erl, 1lkk A,1ctj, 1bpi, 1arb, 1igd, 1ifc, 7rsa, 1cus, 1cse I,1cse E,1ptx, 1aac, 1plc, 1xso A,1ycc, 1rcf, 3ebx, 1rro, 4ptp, 1fus, 135l, 3sdh B,2sga, 2hbg, 2ctc, 256b A,1poa, 2end, 5p21, 2olb A,1xyz B,1eca, 2phy, 1mbd, 3b5c, 1xnb, 2mcm, 2cba, 4xis, 1flp, 4gcr, 1isu B,1tca, 1cka A,2er7 E,1ezm, 2ayh, 3pte, 2rhe, 153l, 1mla, 1tgx C,1whi, 1ptf, 2rn2, 1ppn, 1hfc, 4fgf, 1thb A,1paz, 1csh, 8tln E,2ovo, 3grs, 2sil, 1gmp A,1abe, 3chy, 1lid, 1lam, 1php, 1mrj, 1nif, 1nfp, 1snc, 1scs, 1arv, 1gai, 1fnc, 1dad, 1vsd, 1smd, 3dfr, 1htr B,1htr P,451c, 4icb, 4dfr B,1cnv, 1kap P,1phd, 2cpl, 3lzm, 1fxd, 1edg, 2mhr, 2gdm, 1knb, 2trx A,1sri B,1mba, 2dri, 1fkj, 2wrp R,1ads, 1frd, 1cpc A,1cpc L,1tta B,2utg B,1aky, 3cla, 2msb A,1ctf, 1mol B,1cot, 1gof, 2ltn C,2ltn B,2ccy B,1sbp, 1onc, 1wfa A,1nsc B,1thv, 1rop A,1din, 2bbk J,2bbk L,1iae, 1osa, 8dfr, 1pdo, 1ilk, 1nnc, 3cox, 1gca, 2aza B,1thg, 1kpt B,1vhh, 1nar, 1atl A,2gst B,1amp, 2cyr, 3tgl, 1ida A,1hbq, 1hpi, 1btl, 1mzm, 1bhp, 1ubi, 1lst, 2pii, 1chd, 2bop A,1pgs, 2cyp, 1ido, 1axn, 1hpm, 2ohx B,2tgi, 8fab D,1lau E,1pda, 9wga A,1pk4, 1tad B,1hxn, 1brn M,2sic I,1gd1 R,1mng A,2nad A,4enl, 1byb, 1hyl B,1dup A,1ept A,1tml, 1ept B,1cel A,1muc A,1glq A,1tph 2,1bbh A,2por, 1tgs I,1hyp, 1bmd B,1bdo, 1lmb 3,1pbe, 1gpr, 1tif, 1bgh, 2mnr, 1cmb B,1npk, 1ten, 2hts, 1ede, 1mml, 7aat B,1slt A,1ytb A,2abk, 1vca B,2spc A,2fx2, 1flr L,1hgx B,2lhb, 2rsp A,2cmd, 1fba A,1reg Y,1fiv A,1svb, 1ova C,1rsy, 2prd, 1lis, 1abo A,1cns A,1pnk B,1pnk A,1trk A,1rec, 2ebn, 2chs G,1dvf D,1doi, 1mpp, 1oac B,1rva B,1ukz, 1prn, 1mka A,1rcp B,1daa A,1pne, 1lts E,1lts A,1lts C,1cgt, 2hpd B,1otf A,1dpe, 1dsb B,1aoz A,1oyc, 2tsc A,1puc, 1esf A,1clc, 1gp1 B,2i1b, 1gky, 1pii, 1bam, 2dnj A,1wht A, 1wht B,1alk A,1igs, 6ins E,1acf, 2scp B,1led, 2psp B,1ris, 1lct, 2kau B, 2kau C, 2kau A, 1nba D, 1lki, 2ora, 2pia, 1rcd, 1kpb A,3il8, 1mjc, 1gsa, 1gox, 1frp A, 1fia B, 1poc, 1cdc B,1r69, 1wdc A,3rub S,3rub L,1vid, 1wdc B,1ddt, 1sra, 1cdk B,2pgd, 1ade B, 1hoe, 1pbn, 1bbp B, 1ecp B, 1cfb, 6ldh, 1btn, 1csn, 1tig |
| List of the hits with |
|
| in SCAN3Da: | GPF(28) 1gmp A, GPY(56) 1fus, PPF(10) 1tgx C, SPF(404) 1thg, YPF(4) 1mng A |
| in CSD: | PVANSB, PVANSB, ANTAML10 |
| List of the hits with |
|
| in SCAN3Da: | APY(72) 1lct, APF(284) 2olb A, APF(17) 1lst, MPF(367) 1pnk B, PPY(39) 2erl, PPF(24) 1nif, SPY(133) 1rcd, YPY(237) 1ade B, VPF(251) 3tgl |
| in CSD: | ALPRAL10, ANTAHC10, DUTLAF10, DUTLAF10, GIPKAR10, JUJHUR, JUXHAL, LACSUD, LINCOA, PAANTD, PAANTD01, PVANTS, SUMNOD, VEDJIX, VOWZUC |
| a The three first letters give the peptide sequence; the number in parentheses is the residue number of the aromatic residue; this is followed by the PDB 4-letter code and the subunit identifier. | |
| 1H- and 13C NMR chemical shifts [ppm]b of Ala-Pro-Tyr measured at T = 273 K | ||||||||||
| Res. |
|
|
|
|
or H |
|
|
|
or H |
|
| trans | ||||||||||
| Ala |
|
|
|
|
|
|
||||
| Pro |
|
|
|
|
|
|
|
|
|
|
| Tyr |
|
|
|
|
|
|
|
|
||
| cis | ||||||||||
| Ala |
|
|
|
|
|
|
||||
| Pro |
|
|
|
|
|
|
|
|
|
|
| Tyr |
|
|
|
|
|
|
|
|
||
| a The values characteristic of the
cis-transproline(i-1)-aromatic(i) interaction are
given in bold.
b The chemical shifts are given with respect to the DSS reference compound (Wishart and Sykes, 1994). |
||||||||||
| 1H-1H J-coupling constants [Hz] of Ala-Pro-Tyr measured at T = 273 K | ||||||||||||||||
| trans | ||||||||||||||||
| Ala |
|
5.9
|
|
6.9
|
||||||||||||
| Pro |
|
5.8
|
|
8.3
|
|
-
|
|
7.0
|
|
7.0
|
||||||
|
|
7.0
|
|
7.0
|
|
-
|
|
-
|
|
-
|
|||||||
|
|
10.40-
|
|
10.40-
|
|
-
|
|||||||||||
| Tyr |
|
6.8
|
|
4a
|
|
8a
|
|
-14a
|
||||||||
| cis | ||||||||||||||||
| Ala |
|
5.2
|
|
6.9
|
||||||||||||
| Pro |
|
2.3b
|
|
8.2b
|
|
-12.6
|
|
6.7
|
|
1.4
|
||||||
|
|
11.6
|
|
7.3
|
|
-12.6
|
|
8.9
|
|
9.8
|
|||||||
|
|
2.5
|
|
7.4
|
|
-12.0
|
|||||||||||
| Tyr |
|
7.2
|
|
11.4
|
|
4.7
|
|
-13.9
|
||||||||
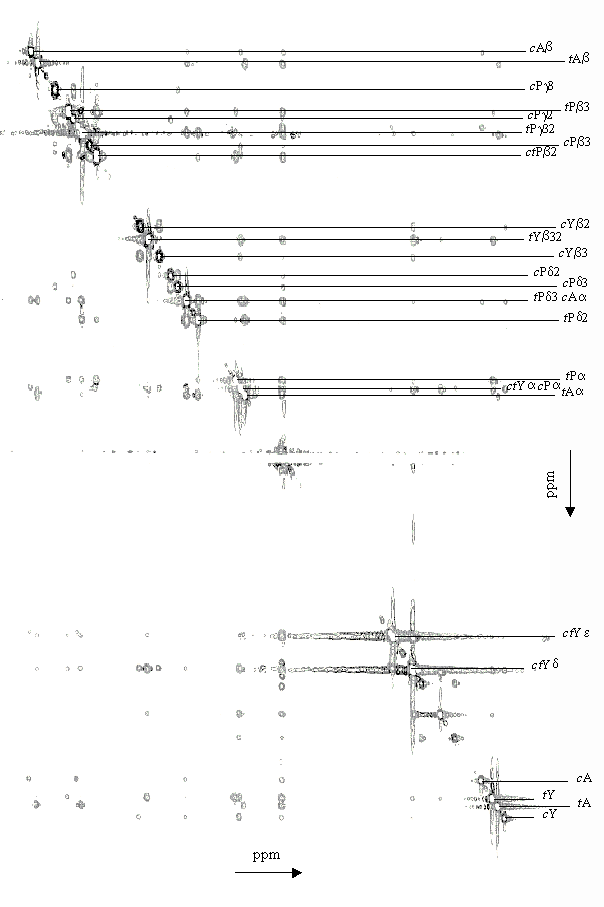
| List of the inter residue ROESY peaks c of Ala-Pro-Tyr measured at T = 273 K | ||||||||||||||||
| trans |
|
|
|
|
|
|
|
|||||||||
|
|
||||||||||||||||
| cis |
|
|
|
|
|
|
||||||||||
| a The 3J-coupling of
Tyr in the trans form is extracted from the fit between experimental
and simulated 1D spectra.
b The small coupling constant found between H c The inter residue ROESY peaks are given with the one amino acid letter code plus the hydrogen type and position. |
||||||||||||||||